Core B
Neuroimaging and Data Analysis
Todd Parrish, Ph. D. and colleagues analyze all behavioral and neuroimaging data collected at project sites to examine models of language recovery. The Neuroimaging data include structural (anatomic and diffusion), functional (task-based and resting state), and physiologic (perfusion). State-of-the-art data processing machinery is used to automatically analyze the data and add them to the project- wide database. These data are used to address specific questions about how the brain recovers from aphasia and the biomarkers that predict language recovery.
Dr. Parrish is a Professor in the Departments of Radiology and Biomedical Engineering at Northwestern University and he directs the Neuroimaging core at Northwestern University’s research imaging facility. His background is in biomedical engineering and biophysics related to MRI methods. He is an expert and leading authority on MR based neuroimaging methods and analysis.
Recent Projects
Northwestern University Neuroimaging Data Archive – NUNDA.
NUNDA is a resource for managing study data collected by the Northwestern University neuroimaging community. It includes a secure database, automated pipelines for processing managed data, and tools for exploring and accessing the data; all while having the advantage of using Northwestern’s high-performance computing cluster (QUEST) for the processing of the image data. Core B has successfully implemented the NUNDA database system allowing its access to users authorized by the specific study’s investigators. As of October 1st, 2018, NUNDA has grown to 291 projects, 14,049 subjects, and 21,537 imaging sessions. The workflow of NUNDA includes: Acquisition → Archive → Automated Processing → Multi-modal Integration → Exploration & Discovery. Image files are transferred via the internet to NUNDA after it has been de-identified. Automated image processing routines then calculate quality assurance metrics that are shared with the site submitting the data. Once approved, the data enters the general database and is automatically processed. Summary images and intermediate data are stored in the database and quantitative region of interest (ROI)-based summary measures are stored in a spreadsheet that is integrated with the participant’s demographic, clinical, and neurobiological measures.
Processing & Analysis. Processing and analysis for NUNDA is done by using an open-source pipeline tool called, PipelineRunner, to direct and monitor automated image processing routines. The processing pipelines utilize software packages used in the neuroimaging field and a pipeline processing menu allows the user to select specific parameter values for all steps or read them from a configuration file. Additionally, by using QUEST, all subjects can be processed in the same time it takes to process a single subject.
Harmonizing imaging protocols across sites. We have achieved the harmonization of the multi-modal data acquisition across the imaging platforms at Northwestern University, Harvard University (Massachusetts General Hospital) and Johns Hopkins University. The Boston and Northwestern sites have the exact same hardware (Siemens Prisma) running at the same software level, which makes harmonizing very straight forward. The Hopkins site however, has a Philips Achieva 3T system with imaging experts to support protocol translation. Core B worked closely with Hopkins to make sure that the protocols matched. All of the magnets being used in this project are dedicated to research activities, which allows the investigators to control software and hardware upgrades, insuring scanner comparability across sites for the duration of the project.
Quality Assurance monitoring and measures. Quality assurance (QA) of imaging data is performed on the participant’s data as soon as it is uploaded into NUNDA. Signal intensity, temporal signal to noise, background noise, percent standard deviation, percent peak to peak, ghost to noise, as well as the Weisskoff noise estimate is calculated from the data to assess the quality of data. QA for the diffusion data is based on motion parameters estimated from the data, the signal to noise of the b=0 scans, and the number of directions that are corrupted by motion. Whereas, anatomic imaging data (T1 and FLAIR) utilize a brain masked signal to noise measure, and a measure of the background noise in the phase encode and readout directions to estimate motion. Finally, the ASL data utilize overall static signal to noise ratio (SNR) and temporal SNR of the time series data once outliers have been removed. The QA parameters for each participant are automatically calculated and swept into a master database and spreadsheet. Core B monitors these metrics and identifies participants that have abnormally low QA measures. The site is notified so they can work with the imaging center to determine the origin of the poor-quality data. Once the service issue has been resolved or if it is participant related (instructed to not move), the participant’s scan will be repeated.
Structural MRI data processing and analysis. We use a two-transform method to take neuroimaging data to Montreal Neurological Institute (MNI) template space. As an example, BOLD data is rigidly aligned to the native structural scan and from there the native structural scan is transformed to MNI space through a concatenated linear and non-linear transformation. Core B has employed a virtual brain transplant method that automatically identifies the lesion, and uses this information to “copy/transplant” brain from the normal hemisphere into the lesion region, then applies the non-linear DARTEL warp to the transplanted brain. The subsequent nonlinear warp is then applied to the native brain to be put into MNI space. The transplant method hinges on the proper identification of the lesion. In order to improve the detection of the lesion, we have recently developed a deep learning network that was trained using 50 different brains (from different sites) and then tested on the remaining subjects, resulting in a Dice index of 0.82. Therefore, currently, lesion maps no longer require to be heavily manually edited.
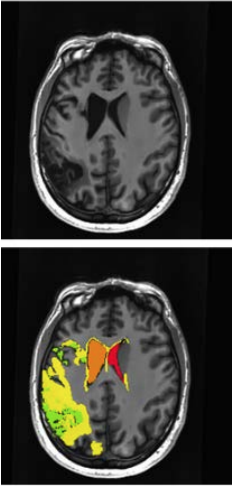
Geographical perilesional definition: Geography based perilesional masks are created by dilating the lesion mask by a specified distance and then subtracting out the lesion, creating perilesional rings of a specific distance. The application of grey and white matter masks, a hemispheric mask (left), and a whole brain mask, is able to limit the perilesional mask to the left brain. This mask is used to test longitudinal changes in the perilesional zone. Using multiple dilation values allows one to test geographic rings for support of language recovery. A separate set of hyperintensity-based lesion masks were created based on the hyperintensities detected in the FLAIR data. In chronic stroke, the major portion of the lesion mimics cerebrospinal fluid (CSF) signal intensity (Figure 1, green) which can be identified on the MPRAGE scan (method of segmentation described above); however, FLAIR imaging is sensitive to hyperintensities that represent gliosis (Figure 1, yellow). These additional lesions can provide insight on brain regions required for language recovery. Furthermore, the presence of hyperintensities in the perilesional region provides information about what regions are capable of recovery.
Perfusion analysis. Perfusion-weighted images from the pCASL scan were pre-processed using a pipeline incorporating Statistical Parametric Mapping (SPM8, Wellcome Trust Center for Neuroimaging, London,UK), and code developed in-house with Matlab R2013a (Mathworks, Natwick, MA) and implemented on the Northwestern University Neuroimaging Data Archive (NUNDA). These methods were used to examine perfusion in the right and left hemispheres of 35 aphasic and 16 healthy control participants. First, the raw EPI images were briefly aligned to the first image of the time-series in order to extract 6 motion-related measures for the timeseries. The motion parameters and global signal were regressed out of the time-series to remove motion-related and physiological fluctuations in the signal. The image time-series were then denoised using an optimized non-linear-means filter. Perfusion-weighted time-series were also generated using pairwise subtraction, and outliers were removed based on previously published guidelines. Time-averaged perfusion maps were then mapped to MNI template space based on the transformations calculated from the high resolution T1 images of the same session.
Physiology based definition of perilesional: The goal of this project is to develop a physiologically based definition of the perilesional space. Core B developed a Gamma based hidden Markov random field model to identify the hypo-perfused tissues that are believed to be perilesional, Figure 2. The method maps the 3D perfusion data to a higher order space where it is fit using gamma distributions (instead of a Gaussian distribution, since the perfusion is not symmetric about zero). Using a numerical methods solution for the expectation maximization, one is able to obtain the mean perfusion for each gamma component. The perilesional (hypo- perfused region) is assumed to be the lower mean value. Therefore, a hypoperfused map is created by labeling these voxels (Figure 2). As a result, the physiologically derived perilesional mask is used to determine BOLD changes related to domain specific tasks. The measures are incorporated into the participant’s database record and are then used in the statistical models of treatment and recovery.
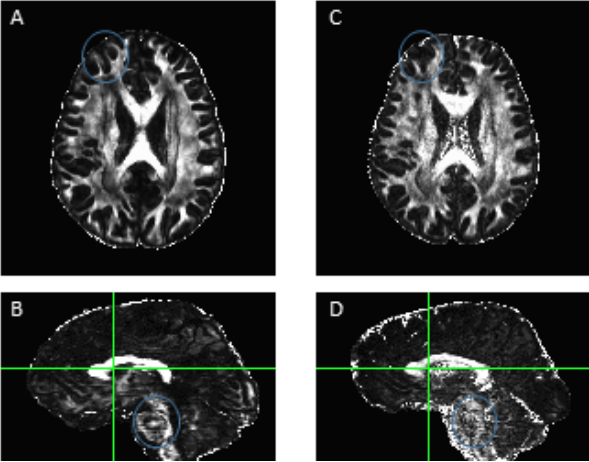
Diffusion image processing and analysis. We developed our own advanced diffusion processing pipeline (ADPP) that follows similar steps to the diffusion pipeline data in the FSL software package (FMRIB, Oxford University) but allows greater spatial mapping in order to maintain the resolution of the acquired data. Image processing methods were implemented primarily using AFNI and Camino (http://afni.nimh.nih.gov/afni), (http://camino.cs.ucl.ac.uk). Processing consists of multiple steps that obtained orientation dispersion metrics for grey matter and “free-water” estimates of the fractional anisotropy (FA) in both grey and white matter. ROIs (e.g., regions within spoken naming, orthographic, and sentence processing networks) are used to extract the DTI indices of white matter integrity and then each ROI is masked to include only white matter. The resultant, ROI-specific DTI metrics, are then stored in the database and compared between time points. Whole brain analyses are also conducted due to the higher quality of our diffusion data and Figure 2: Example of the hypo- (blue) and normal (yellow) perfused regions in two different stroke participants. The lesion mask generated by our automatic segmentation algorithm is shown in red. An additional left hemisphere mask is used to limit analysis of the hypo-perfused tissue. Figure 3: ADPP results in A and B, FSL output results in C and D. We have developed a pipeline that calculates the Neurite Orientation Dispersion and Density Imaging (NODDI) metrics, which provides an estimate of the FA without the contamination of “free water” in each voxel. Using our improved diffusion pipeline, the FA maps are much sharper and allow for better detection of the skeleton. Probabilistic DTI fiber tracking is also conducted using the probabilistic diffusion tractography software, which accounts for crossing fibers. The ROIs masked for white matter will serve as seed and waypoints to examine language network pathways.
Functional task-based data processing and analysis. The preprocessing of fMRI data is performed with the RobustfMRIpreprocessing pipeline that we have developed. The processing steps include, realignment, co-registration to native T1 using rigid and brain boundary registration (BBR), spatial smoothing, and normalization to the Montreal Neurological Institute (MNI) brain template using the non-linear warp from the T1 volume processing. Data censoring is performed by setting FD (Framewise Displacement) and DVARS (volume-to-volume variance) thresholds. Our data censor creates a regressor that includes the time points including the motion and 2 subsequent volumes in order to reduce the effects of motion – following Power’s recommendation. Since there is no deletion of the data, the same task vectors can be used. All task data is then pre-processed and saved in NIFTI format to our database. Analysis for all tasks for all subprojects in the P50 are conducted by the Neuroimaging and Data Analysis Core and the results (beta estimates) are entered into the participant’s database record. Finally, BOLD signal change maps are estimated for individual participants according to a fixed-effects general linear model, with regressors formed from the task events combined with a canonical hemodynamic response function.

Resting state analysis: As stated above, we developed the RobustfMRIpreprocessing pipeline which conducts preprocessing and data analysis (ROI to ROI correlation, ROI to whole brain correlation, amplitude of low frequency fluctuations (ALFF), ReHo, and total connectivity). The preprocessing steps include motion correction, spatial smoothing, linear trend removal, and temporal band-pass filtering of the data. However, before we conduct the correlation analysis, several sources of spurious variance are removed from the data through linear regression: (i) Framewise Displacement (FD), (ii) Volume-to-volume variance (DVARS), (iii) signal from the CSF, and (iv) signal from white matter. The RobustfMRIpreprocessing pipeline can accept ROIs defined in a number of different ways within the same analysis. ROIs can be defined as, spheres of a specified size at a specific coordinate in MNI space, as a binary mask image, or defined as a particular ROI in an atlas such as the Harvard/Oxford, Desikan, Destrieux, or AAL atlas. The ROIs are then masked by the participant’s grey matter map. Next, the mean time course of each ROI is correlated with the mean time course of the other ROIs to form a correlation matrix of all the ROIs. In addition, ROI to whole maps are calculated, where the mean ROI time course is correlated with each voxel in the resting state data. Individual correlation coefficients are then converted to z-scores (“connectivity score”) by using Fisher’s r-to-z transformation to improve the normality of the analysis. The ROI to ROI results are stored in the database to be used in the group analysis of treatment effects, connectomic analysis, and predictors of language/brain recovery. We have also employed voxel-wise statistical models of the fALFF signal (fractional amplitude of low frequency fluctuations, a generalized brain activity measures that is normalized within subject) to understand the meaning of differences in activation level. Our major analysis goals are to characterize the neurotopography of distribution of fALFF values across the entire brain and also more specifically for the brain regions involved in key cognitive networks (default mode, attention, sensorimotor, orthographic, spoken naming, sentence processing). Analyses evaluates how these distributions: (a) compare to those of healthy, age-matched controls, (b) are similar/different across deficit types, (c) correlate with key language and cognitive variables, and (d) change as a result of treatment.
Predicting Post-Treatment Recovery: The group ICA data driven approach is used to investigate how baseline visit resting state fALFF (fractional amplitude of low frequency fluctuations), predicts the participant’s recovery for each deficit type. Group ICA Of fMRI Toolbox (GIFT) is used to identify 20 spatially independent networks using all of the baseline and post treatment data. The networks are back-projected to the native brain space, where the average time series is created and used to generate a fALFF value. The baseline fALFF is then included in a liner regression model to predict the domain specific recovery score. When the fALFF value is combined with the behavioral data the model becomes more accurate and has smaller 95% confidence intervals, Figure 4.